The air-water interface (AWI) is essentially the chaotic boundary where a solution meets the air. When protein molecules in solution contact the AWI, they tend to align themselves along the interface with the hydrophilic regions facing inward towards the water environment and the hydrophobic regions facing outwards towards the air environment.
In cryo-EM sample preparation, a small volume of purified protein solution is deposited onto a 3-mm grid forming a high surface-to-volume droplet with a thickness typically less than 100 nm. Ideally, the particles remain randomly oriented and suspended in the liquid layer prior to vitrification. However, a significant number of protein molecules diffuse to the air-water interface in less than one millisecond [1]. This could result in the molecules aligning themselves in a preferred orientation meaning the desired random distribution is skewed [2]. Also, the protein molecules along the air-water interface could denature or unfold, losing their native structure. That leaves a large part of the sample vulnerable at the interface before being vitrified. All of these could impact the quality and resolution of the 3D reconstruction that results from the vitrified samples.
The AWI presents one of the most difficult problems in cryo-EM sample prep. Numerous solutions, including the use of detergents and custom-made grids, are being explored to overcome the deleterious effects of the AWI and have the macromolecule randomly oriented while also maintaining the native structure.
Can speed mitigate the effects of the air-water interface?
One of the simplest approaches that are being investigated to mitigate the effects of AWI is to speed up the sample preparation process, thereby reducing the time between sample deposition and freezing. But can the effects of the AWI be outrun? What would be the shortest exposure needed to outrun this “deadly touch”?
To understand this better, let’s explore the timescales for protein diffusion. If we consider a protein’s free movement across a cell, the diffusion coefficient tells us that it takes about 10 milliseconds to travel across a 1-µm E.coli bacterial cell [3]. Comparatively, a sample layer in a cryo-EM grid used for single particle analysis (SPA) could be as thin as 50 nm, consisting mostly of water wherein molecules diffuse about 10x faster. That means that the same protein could migrate across the thickness of a typical sample layer within 50 microseconds. It has also been calculated that a 100-kDa molecule will have 1000 interactions with the interface of 100 nanometers in a second [4]. Considering these timescales, it is difficult to imagine mechanics that can immobilize all particles before their Brownian motion leads them to the AWI. Unless the liquid is shielded from the surrounding air altogether, it’s unlikely that the sample preparation process can be fast enough to outrun the effects of AWI.

What else can be done to reduce AWI effects?
The addition of detergents and additives to the sample-containing solution is another way in which macromolecules can be shielded from the AWI. However, it can be difficult in practice to optimize the additive chemistry during this important step due to the inherent inconsistencies in the traditional blotting process used to remove the excess liquid from the EM grid during sample preparation.
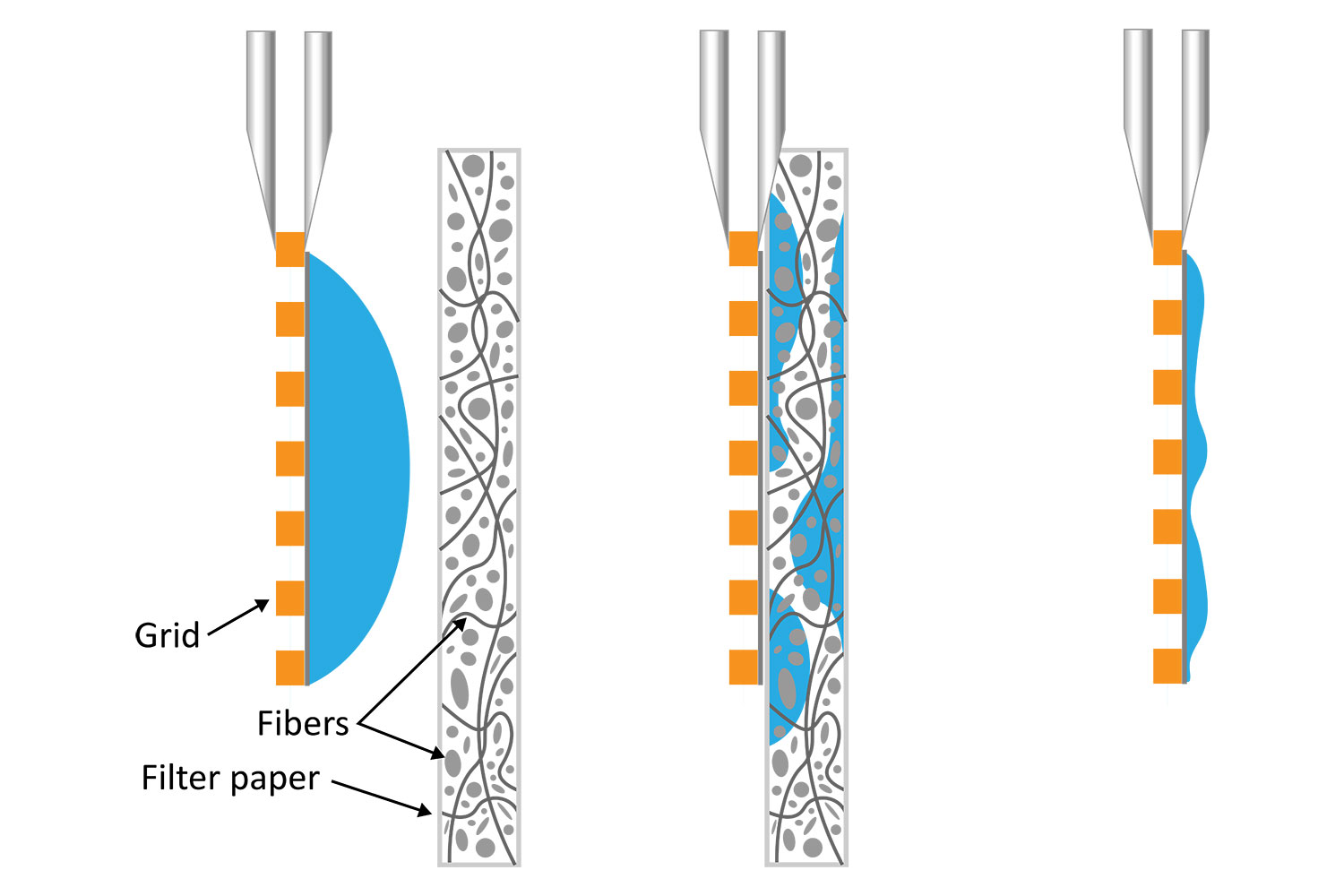
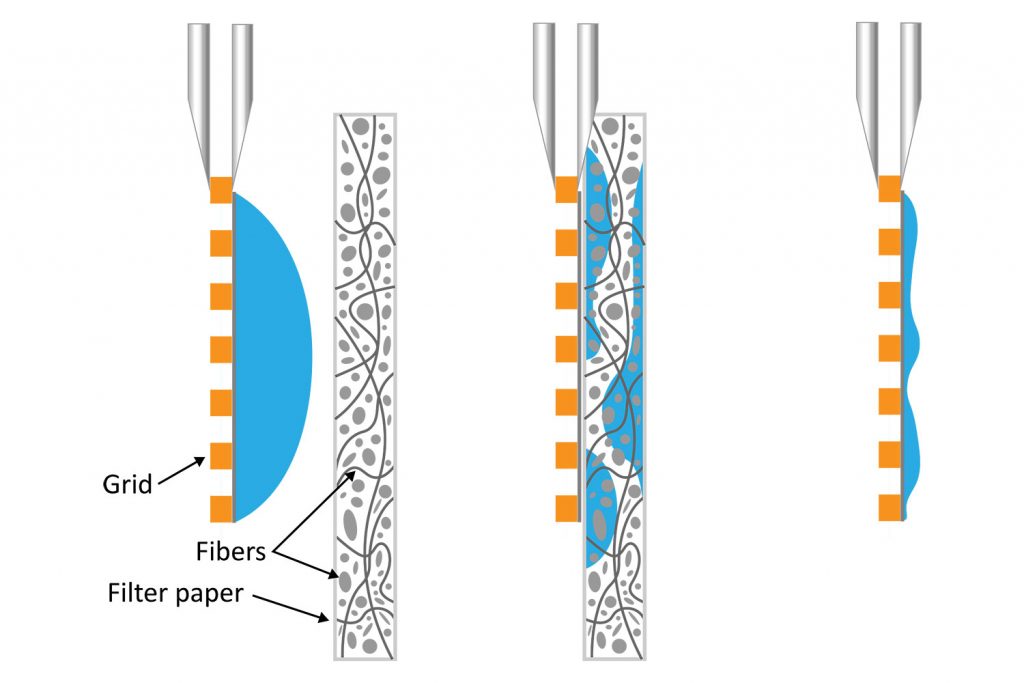
It is well known that blotting results in poor ice layer thickness reproducibility from sample to sample and within a single EM grid [5]. The deposited solution initially forms a large droplet covering the 3-mm grid surface. Randomly oriented fibers in the filter material induce non-homogeneous capillary forces during blotting, resulting in a non-uniform residual liquid layer upon removal. While commercial tools have been developed to partially automate the deposition and blotting conditions, the fundamental randomness in the microscale fluidic behavior that leads to sample layer inconsistency, and hence varying AWI interactions, cannot be eliminated. This ultimately makes it challenging for researchers to evaluate the effect of detergents or additives on their cryo-EM sample preparation.
Automating Cryo-EM sample preparation
The success rate of grid preparation is considered low, with an average of 10 – 50% of grids considered data collection worthy even after having optimized sample conditions. This reinforces the idea that control throughout the sample preparation workflow will improve yield and speed-to-data. On the other hand, reducing the variability will make it easier for scientists to optimize the biochemistry of sample solutions and further eliminate AWI effects.
Automating the sample preparation workflow is key to achieving reproducibility and yield of cryo-EM grids. The VitroJet is a commercial instrument that automates and offers control and speed for high throughput of the cryo-EM grid preparation process. Its core technology relies on blotless pin printing to deposit uniform liquid layers from minute sample volumes. Sample layer thickness can be modulated through key protocol parameters including the pin standoff height, pin radius, and pin printing velocity.
The Vitrojet also replaces plunge freezing with jet vitrification, allowing for a more rapid cooling rate and the use of pre-clipped grids. The Vitrojet can also achieve deposition-to-freezing in as little as 80 ms [6]. A highly automated system handling pre-clipped auto grids without manual intervention from the user helps the researchers overcome the inconsistencies of traditional blot-and-plunge workflows and focus primarily on optimizing detergents and additives in cryo-EM sample preparation. This will be imperative as they optimize the conditions to minimize the deleterious effects of the air-water interface.
References
[1] E. D’Imprima, D. Floris, M. Joppe, R. Sánchez, M. Grininger and W. Kühlbrandt, “Protein denaturation at the air-water interface and how to prevent it,” eLife, vol. 8, p. e42747, 2019.
[2] J. Chen, A. J. Noble, J. Y. Kang and S. A. Darst, “Eliminating effects of particle adsorption to the air/water interface in single-particle cryo-electron microscopy: Bacterial RNA polymerase and CHAPSO,” Journal of Structural Biology: X, vol. 1, p. 100005, 2019.
[3] R. P. Ron Milo, Cell Biology By the Numbers, Garland Science, 2015.
[4] K. N. &. C. J. Russo, “Measuring the effects of particle orientation to improve the efficiency of electron cryomicroscopy,” Nature Communications, vol. 8, p. 629, 2017.
[5] M. Armstrong, B.-G. Han, S. Gomez, J. Turner, D. A. Fletcher and R. M. Glaeser, “Microscale Fluid Behavior during Cryo-EM Sample Blotting,” Biophysical Journal, vol. 118, pp. 708-719, 2020.
[6] R. B. G. Ravelli, F. J. T. Nijpels, R. J. M. Henderikx, G. Weissenberger, S. Thewessem, A. Gijsbers, B. W. A. M. M. Beulen and C. L.-I. &. P. J. Peters, “Cryo-EM structures from sub-nl volumes using pin-printing and jet vitrification,” Nature Communications, vol. 11, p. 2563, 2020.