Electron Microscopy in Life Sciences
There are a number of common electron microscopy methods deployed in life sciences for imaging. The basic entry-level is Scanning Electron Microscopy (SEM) used for magnifications beyond optical microscopes, up to 1 million times magnification. Next comes Transmission Electron Microscopy (TEM) which plays a critical role in life sciences by enabling visualization of biological structures at nanometer and sub-nanometer resolution, far beyond the limits of light microscopy.
In life science research, TEM is used to study cellular ultrastructure, organelles, viruses, protein complexes, and macromolecular assemblies, providing detailed insight into structure–function relationships. Scanning Transmission Electron Microscopy (STEM) extends these capabilities by generating high-contrast, nanoscale images whilst also simultaneously enabling high-sensitivity imaging and elemental analyses (viz “EDS” and “EELS”) unlocking complex physical and chemical structures. Cryogenic Electron Microscopy (Cryo-EM) has transformed structural biology by allowing biological samples to be imaged in a near-native, vitrified state without chemical fixation or staining, preserving structural integrity while enabling high-resolution 3D reconstruction of proteins and complexes. Lastly Volume-EM or vEM is a family of techniques whereby materials (in this case cells and tissues) are sliced and imaged to provide a 3D view of the sample. It is often used in conjunction with Cryo-EM. Together, SEM, TEM, STEM, vEM and Cryo-EM form an essential toolkit for advancing molecular and cellular understanding in modern life science research.

What are the common challenges of EM in life sciences?
Despite its powerful capabilities, the use of EM in life sciences presents several common challenges that must be carefully managed to obtain reliable results. These include:
Beam Sensitivity and Radiation Damage:
Biological materials are inherently beam-sensitive, limiting the electron dose that can be applied. Radiation damage can alter fine structural details, thereby reducing image quality and complicating high-resolution analysis, particularly for proteins and soft tissues. Furthermore, the heterogeneous electric properties of organic samples often results in charging artefacts meaning areas that cannot easily be viewed with the electron beam as electrons pool in certain areas.
Throughput and Data Complexity:
High-resolution TEM and especially Cryo-EM generate large datasets that require significant computational resources for processing, alignment, and 3D reconstruction. Data acquisition and analysis can be time-consuming and demand specialized expertise.
Data Acquisition Time:
Data acquisition time is a significant consideration when using TEM, STEM, and Cryo-EM in life science applications. vEM can sometimes take days to acquire let alone process data. Acquiring statistically meaningful datasets using STEM can extend acquisition times to several hours. In Cryo-EM, data acquisition is particularly time-intensive, as high-resolution 3D reconstructions require the collection of tens to hundreds of thousands of particle images under strict low-dose conditions. This process often involves extended microscope sessions and careful optimization of imaging parameters to balance resolution and radiation damage.
How does SenseAI address the challenges of EM in life sciences?
SenseAI has developed a compressed sensing approach that integrates intelligent subsampling with blind dictionary-learning–based inpainting to significantly reduce the number of electrons (or indeed any beam) required to generate a full-resolution image (Figure 1). Rather than acquiring a complete dataset, the system deliberately captures only a sparse subset of pixels or probe positions, often as little as 1 – 10% of the total data, and then accurately reconstructs the full image through advanced computational algorithms.
Importantly, SenseAI reconstructions are mathematically validated and constrained by the acquired data, ensuring that the algorithms cannot “hallucinate” or introduce false structural information, a common concern with many AI-based methods. As a result, the reconstructed image faithfully represents the true structure of the specimen. In some cases, image quality can exceed that of a conventionally fully sampled dataset, as reduced electron or photon dose minimizes beam-induced damage, while the faster nature of subsampling can also reduce motion-related blurring. Note that for TEM the benefits are similar albeit the application is different: TEM acts like a ‘planar’ beam (similar to an x-ray at the dentist) rather than a scanned beam. So SenseAI is used to ‘denoise’ a low-dose image, rather than subsample the original beam as it isn’t scanned to begin with.
The key benefits of using SenseAI solution are three-fold: it avoids beam damage on delicate samples and charging; it allows faster image acquisition; and it uses up to 100x less data to capture and store those images. By addressing the core physics and practical constraints of using electron microscopy in life sciences, SenseAI makes workflows that were once slow, destructive, or data-limited into fast, efficient, and sample-friendly processes. SenseAI is now working on “4D STEM” which is an ultimate STEM method involving rich data gathered at each point. It is using this to transform one of the key ‘workhorse’ methodologies of the life sciences industry – Single Particle Analysis or SPA. This is a methodology for determining the precise structures of complex organic molecules and is typically highly involved and laborious process collecting data from many millions of sample particles. The benefits of SenseAI are to achieve a far greater resolution in less data meaning a multi-order of magnitude reduction in time.
How is the SenseAI solution being implemented?
Several research institutes across the globe have already implemented SenseAI on their electron microscopes and are seeing significant improvements in data acquisition and analysis workflows. Here we elaborate on the use of SenseAI at Radboud University Medical Center and King’s College London.
How is SenseAI preserving biological sample integrity at the Radboud University Medical Center?
Radboud University Medical Center, located in Nijmegen, The Netherlands, is one of Europe’s leading academic hospitals and research institutes. Through the Radboud Electron Microscopy Center, founded by Professor Nico Sommerdijk, the institution supports advanced electron microscopy of biological materials, with a strong focus on biomineralization, nanoscale materials, and in situ imaging of dynamic biological processes.
Prof. Sommerdijk’s group specializes in 3D volume imaging, correlative light and electron microscopy, and cryo-electron microscopy. More recently, through the NWO-Groot BIOMATEM project, Radboud has become a national facility for liquid-phase electron microscopy of biological materials, enabling the study of highly beam-sensitive samples under near-physiological conditions.
To address the fundamental challenge of imaging biological materials at low electron dose, the team adopted SenseAI’s compressive sensing approach. Dr. Luco Rutten, a postdoctoral researcher in Professor Sommerdijk’s group, explains: “When sampling biological materials, TEM works by analysing very thin samples, where you need to work with low electron doses so as not to damage the samples. This generates noisier images at lower resolution. At the same time, to study biological process we often need a larger volume to be more physiologically relevant. We use STEM for these thicker samples where we can get a larger depth of focus. In situ is the holy grail for both, particularly when you want a movie of nano-processes and for that you need a very low dose.”
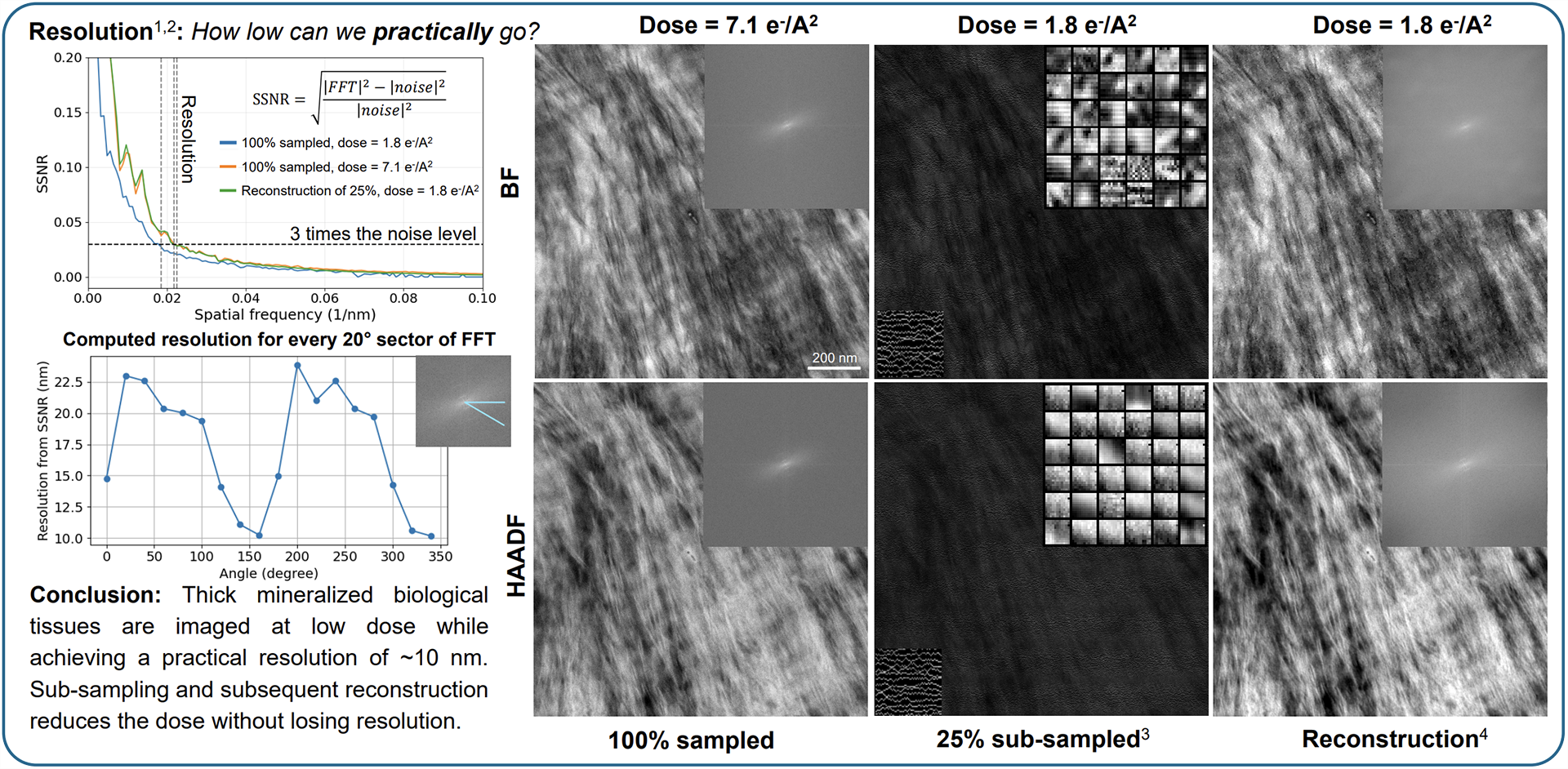
Using SenseAI, the Radboud team has been able to generate high-resolution STEM and TEM images using only a fraction of the electron dose. In studies of collagen fibrils, SenseAI reconstructed images with equivalent resolution to full-dose datasets while using just 25% of the dose, accelerating experiments by a factor of four, illustrated in Figure 2.
Looking ahead, Professor Sommerdijk sees strong potential for SenseAI in even more challenging biological systems, including membrane fusion and neural tissue, stating, “Our goal is to look at cells with electron doses of less than 0.05 electrons per square Angstrom as this is the limit of when enzymes are active. This is where we can use sparse sampling to go to new levels. To reach this limit we need to combine sparse imaging with other strategies to reduce the electron damage e.g. graphene protection. We foresee great potential for the in-situ imaging for biological processes using these methods.”
“We see potential in other techniques where SenseAI can potentially power us forward even more – Cryo FIB-SEM, Raman Spectroscopy, and Confocal Microscopy. SenseAI enabled us to break through technical barriers to take our research to new levels. It is also straightforward and quick to use. We are excited about our next leg of the journey.”
Prof. Nico Sommerdijk – Radboud University Medical Center
How is SenseAI helping accelerate ultrastructural imaging at King’s College London?
King’s College London is home to the Centre for Ultrastructural Imaging (CUI), one of Europe’s most advanced electron microscopy hubs for life science and biological research. The CUI supports a wide user base with expertise in cryogenic electron microscopy techniques, including dual-beam cryo-focused ion beam scanning electron microscopy (cryo FIB-SEM) and cryogenic volume electron microscopy imaging (vEM), which are essential for studying complex, beam-sensitive biological materials.
Cryogenic volume imaging, particularly, presents significant technical challenges. Frozen biological samples are extremely sensitive to electron irradiation, with beam damage and devitrification posing constant risks. At the same time, conventional volume imaging workflows are slow, with acquisition times often extending to 12 – 24 hours or more, severely limiting experimental throughput and flexibility.
Professor Roland Fleck, Director of the CUI, explains the challenge “imaging cryogenically preserved biological materials requires exceptional care, precise control of imaging parameters, and long acquisition times; factors that together restrict both the pace of research and accessibility for new users”.
To address these limitations, the CUI adopted SenseAI’s real-time electron microscopy imaging platform. SenseAI was integrated directly into the CUI’s existing cryo FIB-SEM infrastructure, providing advanced imaging control without requiring additional data science or computer science expertise from users (full interview with Professor Roland Fleck in Figure 3).
With SenseAI, researchers at CUI achieved fast, low-dose, high-resolution imaging of biomaterials while significantly reducing beam-induced damage.1 The platform enabled users to fine-tune imaging parameters for specific materials, improving contrast and accuracy, all through a single, intuitive interface.
Following implementation, the CUI observed a five-fold increase in imaging speed with no loss in data quality. At the same time, electron beam damage and sample devitrification were reduced by approximately 80%, leading to more consistent datasets and fewer imaging artefacts across extended acquisition sessions.
“SenseAI has made a significant impact on CUI’s electron microscopy practice, and we are delighted with the results…The quality of the imaging is superior as well as the speeding up and automation. This has helped us to further our ambitious research, to serve our internal and external users, and move the state-of-the-art forward.”
Professor Roland Fleck – King’s College London
Conclusion
SenseAI’s technology represents a major step forward in electron microscopy, offering a practical solution to one of the field’s most persistent challenges—balancing efficiency, image quality, and data volume. By integrating advanced subsampling algorithms with real-time image reconstruction, SenseAI empowers researchers to capture high-quality data faster and more efficiently, while preserving the integrity of even the most delicate samples. With its compatibility across multiple EM modalities and easy integration, SenseAI is poised to be an essential tool for the next generation of microscopy innovation.
References
- Nicholls, D.; Kobylynska, M.; Broad, Z.; Wells, J.; Robinson, A.; McGrouther, D.; Moshtaghpour, A.; Kirkland, A. I.; Fleck, R. A.; Browning, N. D. The potential of subsampling and inpainting for fast Low-Dose cryo FIB-SEM imaging. Microscopy and Microanalysis 2024, 30 (1), 96–102. https://doi.org/10.1093/micmic/ozae005. ↩︎



