Sample preparation for cryogenic electron microscopy (cryo-EM) can be a demanding and expensive process, requiring several rounds of purification and screening to isolate the macromolecules of interest. Negative stain electron microscopy (EM) is one of the established tools for the initial screening of purified samples, specifically for assessment of particle size, morphology, concentration, and agglomeration. Traditionally, a conventional transmission electron microscope (TEM) is used to image the stained samples. With the introduction of the desktop scanning transmission electron microscope (STEM), an alternate process for sample screening has been developed that is faster and more cost-effective than the traditional methods.
Cryo-EM
Cryogenic electron microscopy (cryo-EM) has emerged as an invaluable tool in the field of structural biology, enabling researchers to visualize vitrified macromolecules and elucidate their three-dimensional structures at near-atomic resolution. During data collection, multiple 2D projection images are acquired at different orientations. These images are then used for 3D reconstruction through advanced computational techniques, ultimately resulting in a high-resolution 3D structure of the macromolecules or complexes of interest at near-atomic resolution.
Cryo-EM is a laborious and time-intensive technique that requires highly specialized skill sets. To get good results, the purified samples must meet a very high standard in terms of quality. So, prior to vitrifying a sample for high-resolution imaging, they are usually screened to ensure that the sample contains the appropriate concentration of particles with the expected sizes and morphologies.

Negative Stain Electron Microscopy
This technique was first demonstrated in the mid-twentieth century by Brenner and Horne who described a protocol for staining virus particles with heavy-metal salts and observing them in a TEM [1]. For particle specimens, the sample is absorbed onto the grid and then a stain is applied to envelope the particles. A high mass-thickness contrast difference is produced by the background relative to the particles in the electron microscope, allowing for a “negative” image of the particles to be visualized at nanometer-scale resolution [2]. The ease of grid preparation and high contrast offer an ideal method for the assessment of particle size, morphology, concentration, and agglomeration [3].
A conventional TEM is generally used to visualize the negatively stained samples. This results in expensive equipment being tied up in screening samples rather than their intended application in high-resolution imaging. The facility costs and frequent maintenance costs can be significant, imposing financial burdens on principal investigators who are relying on it for sample screening. The learning curve on how to use a TEM can be rather steep with new users taking several days to weeks to become proficient enough to operate a TEM independently. Therefore, exploring alternatives to dedicated TEM instruments is an appealing choice for those who want to maximize the benefits of negative stain EM for sample screening.
Desktop STEM
STEM-in-SEM involves using a scanning electron microscope (SEM) to perform transmission imaging. Recently, STEM-in-SEM has been made possible on a desktop system: namely, the Phenom Pharos Desktop SEM with a STEM detector. The powerful new plug-and-play accessory integrates a segmented solid-state detector into a sample holder compatible with standard EM grids and is capable of independent collection of bright-field (BF), annular dark-field (ADF), and high-angle annular dark-field (HAADF) signals.
Thanks to the Pharos’ native resolution, which is better than 1 nanometer, BF-STEM images (analogous to the phase contrast TEM images acquired in traditional negative stain EM workflows) take just under two minutes to acquire once the sample is loaded. Its compact footprint, intuitive software, and resilient hardware offer significant savings in facilities, maintenance, and training costs to its users. Additionally, the Phenom SEM can be programmed to collect data in an automated fashion, allowing for quick inspection of the entire grid. This allows users to collect images containing hundreds of thousands of particles that can be used for statistical analysis.
Using the Phenom Desktop STEM for Negative Stain EM
Some examples of negative stained biological specimens imaged on the new Phenom Pharos Desktop STEM include:
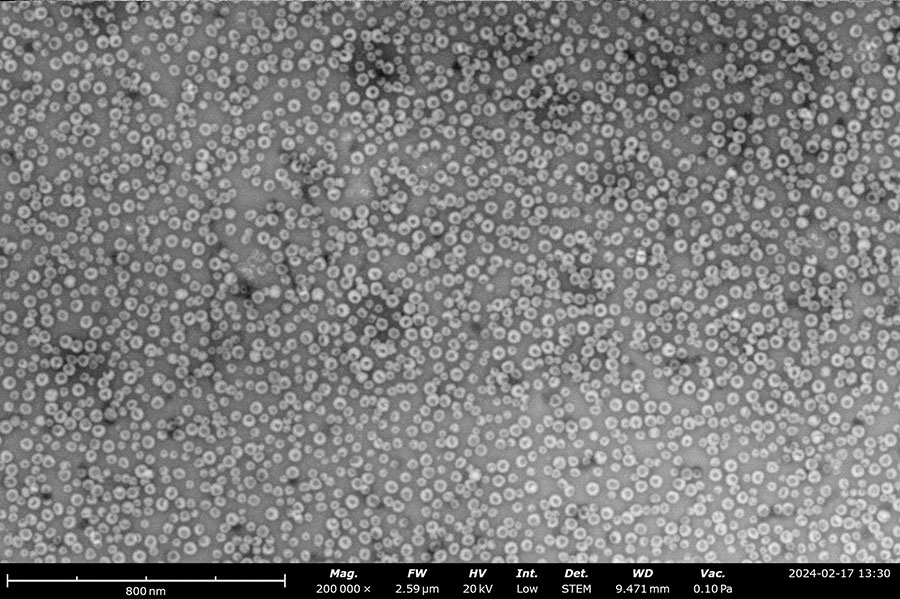
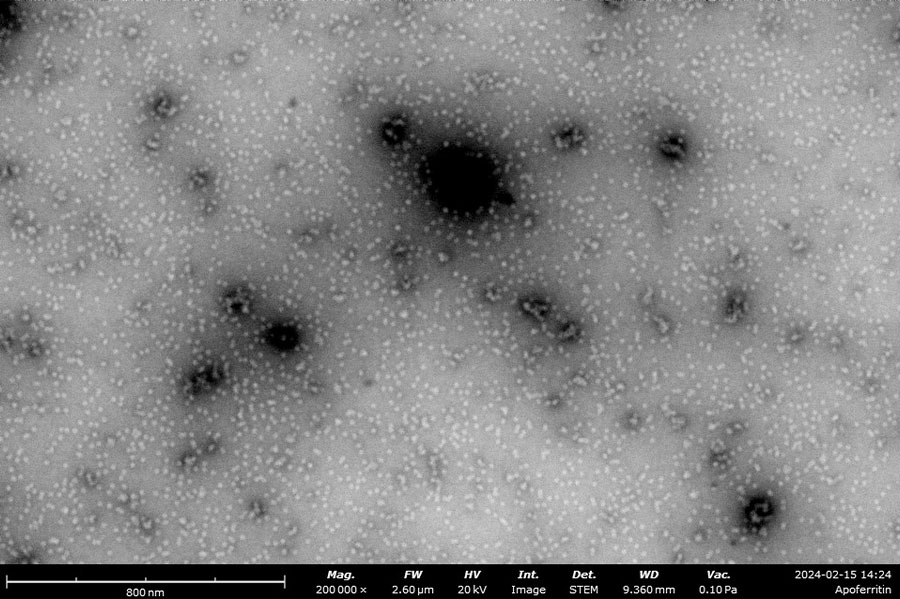


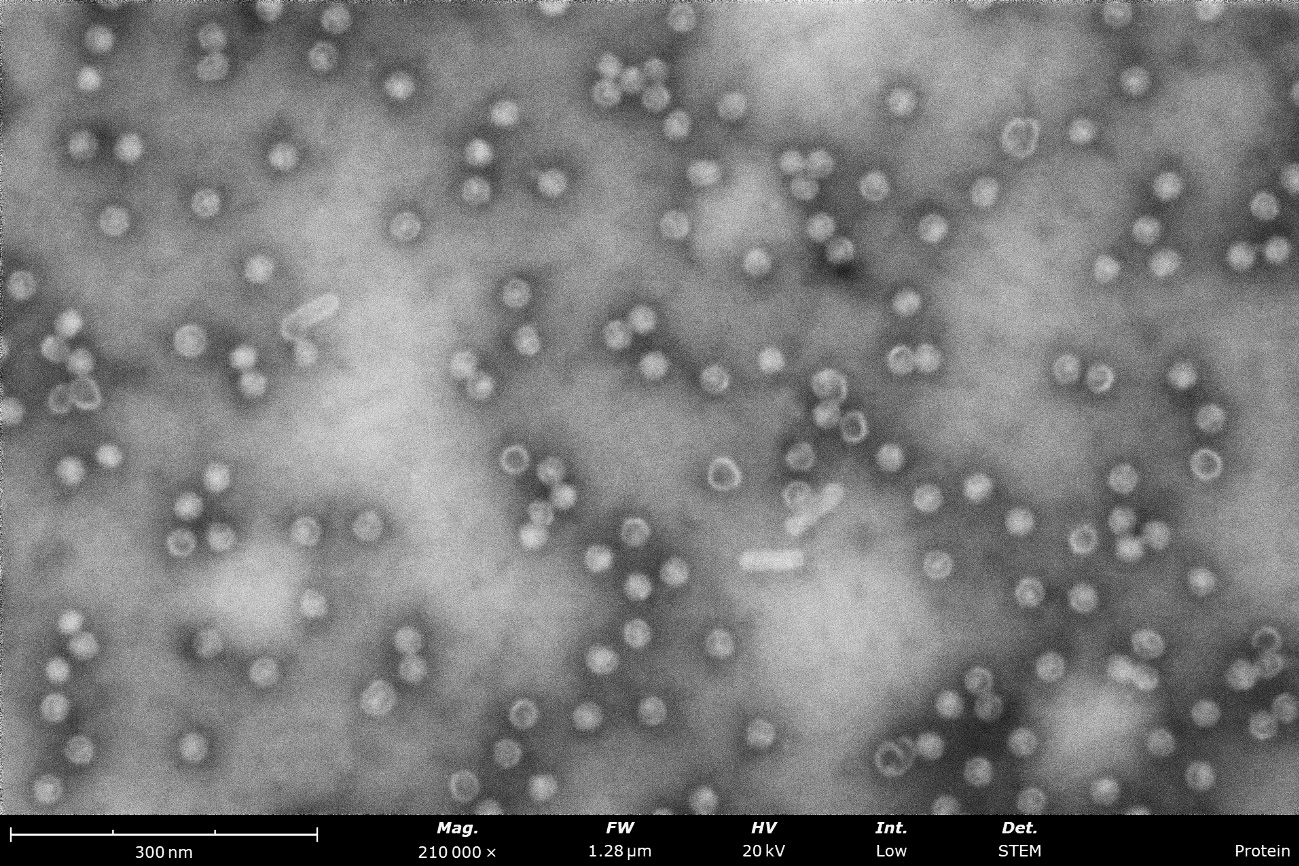
Negatively stained samples of Horse Spleen Apoferritin (left) and Myxococcus Xanthus Encapsulin (right) courtesy of Daija Bobe and Joshua Mendez, NYSBC. Apoferritin particles are measured at approximately 13 nm and Encapsulin is approximately 25-40 nm. These images provide the information needed to screen the effectiveness of the purification process, verify the accuracy of concentration, and identify any agglomeration within the sample.
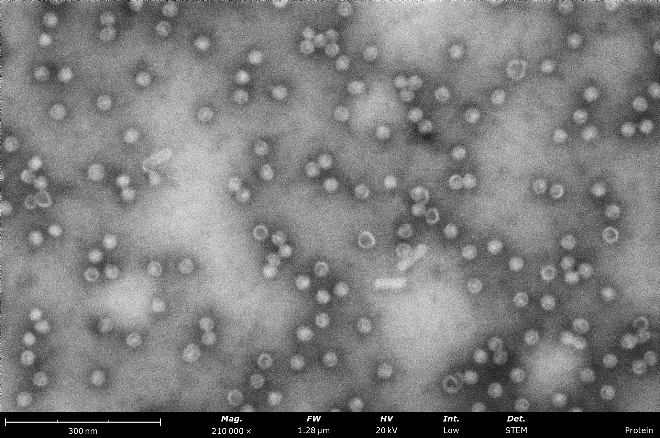

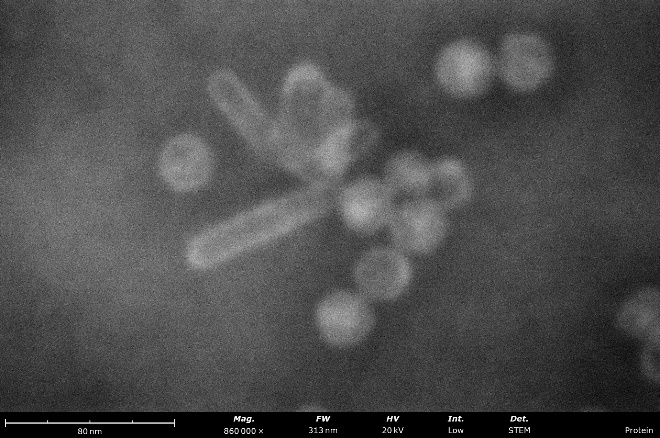
Virus-like-particles stained with nano-tungsten showing rod-like and round morphologies. From these images, a microscopist can measure particle sizes and assess sample purity, agglomeration, and overall distribution to determine if further sample optimization is needed before vitrification for cryo-EM.
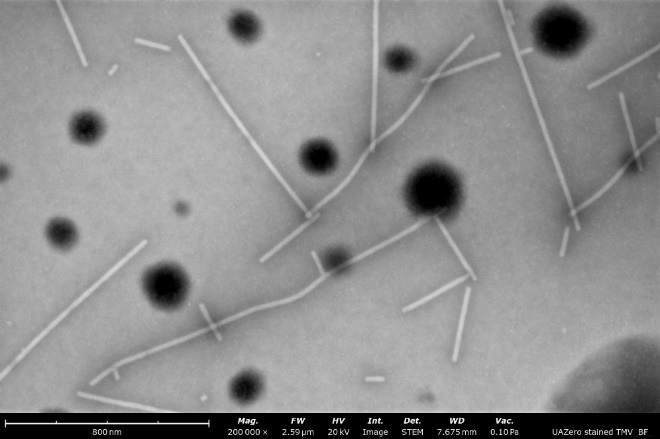
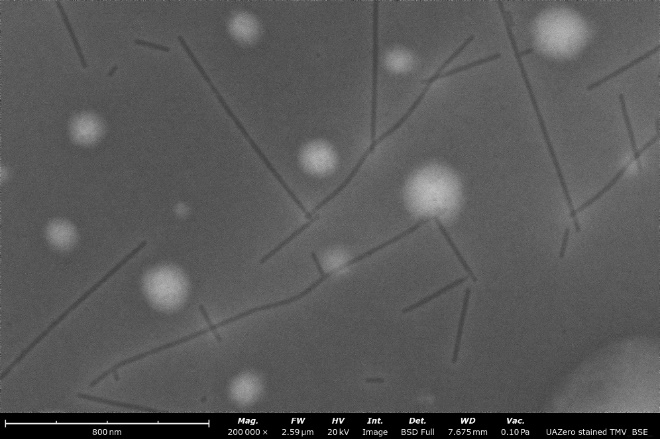


BF-STEM image (left) compared to a backscattered electron detector (BSD) image (right) of a stained sample of tobacco mosaic virus particles. Both images were obtained over the same field of view and notably, the contrast and resolution are enhanced greatly when using the STEM detector.
Summary
For electron microscopists who want a way to do negative staining EM quickly in a cost-effective fashion, the Phenom Pharos Desktop STEM can make that possible. It can also be used to offload work from another microscope or replace a microscope where service support is ending, all at a fraction of the cost. Moreover, as the Phenom functions as an SEM, it provides the capability to inspect additional samples through surface imaging. With its versatility and affordability, the Phenom Pharos Desktop STEM is a valuable addition to any research facility looking to streamline its cryo-EM workflow.
References
[1] S. D. Carlo and J. R. Harris, “Negative staining and Cryo-negative Staining of Macromolecules and Viruses for TEM,” Micron, pp. 117-131, 2011.
[2] J. R. Gallagher, A. J. Kim, N. M. Gulati and A. K. Harris, “Negative‐Stain Transmission Electron Microscopy of Molecular Complexes for Image Analysis by 2D Class Averaging,” Current Protocols in Microbiology, vol. 54, no. 1, p. e90, 2019.
[3] C. A. Scarff, M. J. G. Fuller, R. F. Thompson and M. G. Iadaza, “Variations on Negative Stain Electron Microscopy Methods: Tools for Tackling Challenging Systems,” Journal of Visualized Experiments, vol. 132, p. 57199, 2018.








